5. Inferring Galaxy Parameters (SAN)#
The method described in this page, the Sequential Autoregressive Network, (SAN) is currently the best performing method in the codebase. As a reminder, we wish to estimate the distribution of physical galaxy parameters \(\mathbf{y} \in \mathbb{R}^{9}\), given some photometric observations \(\mathbf{x} \in \mathbb{R}^{8}\); that is \(p(\mathbf{y} \vert \mathbf{x})\).
We draw loose inspiration from other autoregressive models such as [MADE2015], while focusing on the goals of providing fast sampling times (i.e. being able to compute the distribution of \(p(\mathbf{y} \vert \mathbf{x})\) quickly as well as being able to draw samples from it quickly) and maintaining good accuracy; that is, we hope for low test NLL, \(-\sum^N_{i=1} \log p_{\text{SAN}}(\mathbf{y}_{i}\vert \mathbf{x}_{i})\) for pairs of points in the testing dataset \(\mathcal{D}_{\text{test}} = \big\{(\mathbf{x}_{i}, \mathbf{y}_{i})\big\}_{i=1}^{N}\).
5.1. Autoregressive Models for Distribution Estimation#
Before considering the full details of the SAN model, we first review autoregressive models.
Strictly speaking, an ‘autoregressive model’ is one where the output of the model (usually indexed by time) depends on the previously outputted values of the model and some stochastic term. For example, they are commonly employed to predict the next value(s) in a time series.
In this context however, we use a model with this autoregressive property to estimate a multivariate distribution, one dimension at a time. That is, rather than specifying the value of a stochastic process at a given timestep (or index), each iteration of the autoregressive model outputs the next dimension of the multivariate distribution.
Usually, when estimating / learning distributions over data, we must be careful to make sure that the distribution normalises. For example, if we observe a set of points \(\{\mathbf{x}_{i}\}_{i=1}^{N} \in \mathcal{X} \subseteq\mathbb{R}^d\), and we’d like to discover \(p(\mathbf{x})\), then we must make sure that
Even for small \(d\), this can be an expensive procedure unless this normalisation consideration is explicitly thought about when designing the model.
Autoregressive models side-step the need to compute or even approximate high dimensional integrals by simply factorising the desired multivariate distribution as a product of its nested conditionals:
(On notation, \(\mathbf{x}_{<d} \doteq [x_{1}, \ldots, x_{d-1}]^\top\).)
Recall that during training we have \((\text{photometry}, \text{parameter})\) pairs \((\mathbf{x}, \mathbf{y})\), and that we are not interested in learning the joint \(p(\mathbf{x}, \mathbf{y})\) (as the equation above suggests) so much as the distribution of parameters conditioned on photometry \(p(\mathbf{y} \vert \mathbf{x})\).
We can straightforwardly extend the above to include the conditioning information (now \(\mathbf{x}\)) and instead factorise the data we care about modelling, \(\{\mathbf{y}_{i}\}_{i=1}^{N} \in \mathcal{Y} \subseteq\mathbb{R}^D\), giving us:
That is, so long as we can ensure that the output for the \(d^{\text{th}}\) dimension \(y_{d}\) only depends on the previous dimensions \(\mathbf{y}_{<d}\) as well as the conditioning information \(\mathbf{x}\), then the density \(p(\mathbf{y} \vert \mathbf{x})\) can be efficiently computed as the product of factorised terms. We will refer to this property as the autoregressive property going forward.
For instance, we could compute the negative log likelihood as:
So long as \(D\) remains relatively small (which, for this application, should be no more than about 10), and the individual dimensions are modelled with an easily computed distribution, then the above should remain quick to compute.
5.2. The Sequential Autoregressive Network#
The following shows the architecture of our proposed autoregressive model, which we call a Sequential Autoregressive Network.
Click the image to zoom into the SVG in a new tab if it is too small.#
The network is composed of \(D\) sequential blocks. These are repeated sequences of layers (which do not share weights), which accept as input the conditioning information \(\mathbf{x}\), a set of \(F\) sequence features from the previous block (excluding the first), as well as all the stochastic outputs \(\hat{y}_{d} \sim p(y_{d} \vert \hat{\mathbf{y}}_{<d}, \mathbf{x})\) from previous blocks.
This combination of features is somewhat unusual:
in autoregressive models, the \(d^{\text{th}}\) output usually only depends on the previous outputs \(\hat{y}_{d} \sim p(y_{d} \vert \hat{\mathbf{y}}_{<d})\), but not the ‘sequence features’ or conditioning information \(\mathbf{x}\).
in a recurrent network (e.g. RNN), usually only the ‘sequence features’ (or equivalent) are passed through iterations, and not the network outputs for previous iterations \(\hat{\mathbf{y}}_{<d}\).
Since we can return an arbitrary number of parameters at the output of each sequential block, we are free to parametrise any distribution we like for \(p(y_d \vert \hat{\mathbf{y}}_{<d}, \mathbf{x})\). In the diagram above, we show a Gaussian mixture with \(K\) components, with the \(d^{\text{th}}\) sequential block returning \(K\) locations \(\{\mu_{d,i}\}_{i=1}^{K}\), scales \(\{\sigma^2_{d,i}\}_{i=1}^{K}\), and mixture weights \(\{\varphi_{d,i}\}_{i=1}^{K}\) such that \(\sum_{i=1}^K \varphi_{d,i} = 1\).
This model architecture was found to satisfy the desiderata of fast sampling (one can draw 10,000 posterior samples from \(p(\mathbf{y} \vert \mathbf{x})\) for a single source \(\mathbf{x}\) in the order of 10ms), and reasonable accuracy:
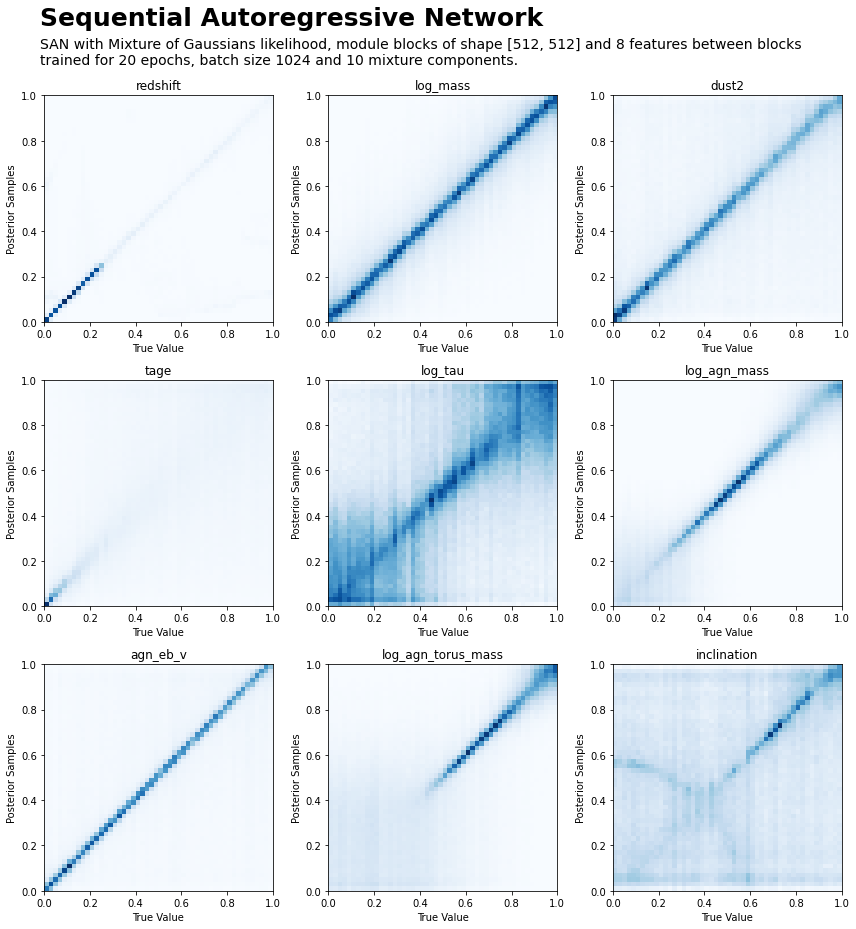
5.2.1. Using the SAN Model#
The configurations for this model (see the inference overview for general configuration information) are defined in
config.py.
- class spt.inference.san.SANParams#
Configuration class for SAN.
This defines some required properties, and additionally performs validation of user-supplied values. See
ModelParamsfor additional configuration values.- Example
>>> class SANParams(san.SANParams): ... epochs: int = 10 ... batch_size: int = 1024 ... dtype: t.dtype = t.float32 ... cond_dim: int = len(ForwardModelParams().filters) ... data_dim: int = len(ForwardModelParams().free_params) ... first_module_shape: list[int] = [1024, 2048, 1024] ... module_shape: list[int] = [1024, 1024] ... sequence_features: int = 16 ... likelihood: Type[san.SAN_Likelihood] = san.TruncatedMoG ... likelihood_kwargs: Optional[dict[str, Any]] = { ... 'lims': t.tensor(ForwardModelParams().free_param_lims(normalised=True)), ... 'K': 10, 'mult_eps': 1e-4, 'abs_eps': 1e-4, 'trunc_eps': 1e-4, ... 'validate_args': False, ... } ... layer_norm: bool = True ... train_rsample: bool = False ... opt_lr: float = 3e-3 ... opt_decay: float = 1e-4
- property first_module_shape: list[int]#
The size of the first module of the network. This is used to build useful initial sequence features, and allows the subsequent blocks to be smaller, reducing memory requirements.
Default: the usual module shape
- abstract property layer_norm: bool#
Whether to use layer norm (batch normalisation) or not
- abstract property likelihood: Type[spt.inference.san.SAN_Likelihood]#
Likelihood to use for each p(y_d | y_<d, x)
- property likelihood_kwargs: Optional[dict[str, Any]]#
Any keyword arguments accepted by likelihood
- property limits: Optional[Type[torch.Tensor]]#
Allows the output samples to be constrained to lie within the specified range.
This method returns a (self.data_dim x 2)-dimensional tensor, with the (normalised) min and max values of each dimension of the output.
- abstract property module_shape: list[int]#
Size of each individual ‘module block’
- property opt_decay: float#
Optimiser weight decay
- property opt_lr: float#
Optimiser learning rate
- abstract property sequence_features: int#
Number of features to carry through for each network block
- property train_rsample: bool#
Whether to stop gradients at the autoregressive step
5.3. References#
- MADE2015
Germain, Mathieu, Karol Gregor, Iain Murray, and Hugo Larochelle. ‘MADE: Masked Autoencoder for Distribution Estimation’. In Proceedings of the 32nd International Conference on Machine Learning, 881–89. PMLR, 2015. https://proceedings.mlr.press/v37/germain15.html.